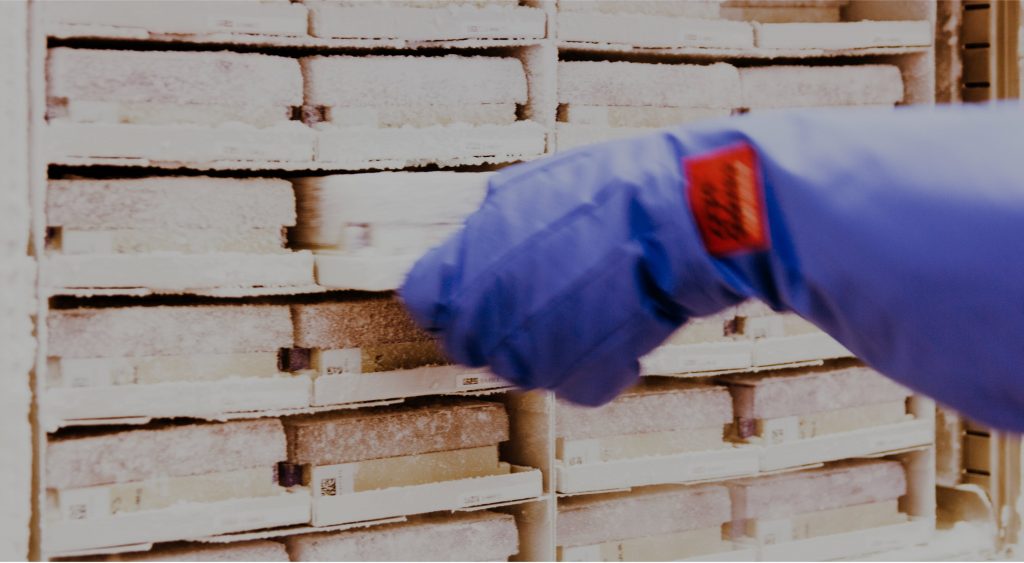
Integrated biobank of Luxembourg (IBBL)
IBBL offers biobanking-related services, including the collection, processing, analysis and storage of biological samples and associated data, in compliance with international quality standards.
Biospecimen research is also performed to optimise sample processing and certify biospecimen quality.
activities
Operations are run under the control of a well-defined Quality Management System (QMS) and certified for the following norms:
ISO 9001:2015 (certification) that specifies the general requirements for quality management systems in client-based organisations .
ISO 17025:2017 (accreditation) that specifies the general requirements for the competence of testing and calibration laboratories for the following methods:
- DNA Quantification by Spectrofluorometry;
- Nucleic Acid Quantification by Spectrophotometry;
- RNA Integrity Measurement;
- 16s rRNA Gene Sequencing.
Finally, operations are also compliant with the principles of ISO 17043:2010 (Proficiency Testing). FIND OUT MORE

Barbieux
Downloads
- The LIH Quality Manual
- IBBL ISO 9001 Certificate
- IBBL ISO 17025 Certificat & Scope
Biospecimen and data generation
Biospecimen Science
- Provide direct research support to the Biorefinery team, other research groups within LIH and future TMOH collaborators to address specific biospecimen-related issues that compromise the quality or efficiency of their work.
- Carry out research that improves the efficiency by which clinical biospecimens are utilised by the research community and generally, enabling more research to be performed with less clinical material.
- Disseminating the expertise of IBBL and LIH by publishing high quality Biospecimen Science Research papers and engaging with the global research community in matters relating to Biospecimen Science.
biorefinery (ISO 9001:2015 certification)
Method Validation and Implementation
- Conducting pre-process/characterisation feasibility and optimisation of new technologies to establish best practices for biospecimen processing
- Formal method validations and implementation of novel methods e.g. cfDNA and Exosomes isolations
- Scientific Equipment Supervision: Implementation, validation and tracing the complete life cycle of scientific equipment
Biospecimen Processing
- Highly standardised and automated processing of biospecimens from continuous routine and static clinical collections
- Highly standardised and automated transformation and derivation of biospecimens
- Highly specialised processing and characterisation e.g. 16S Sequencing, Elispot and ddPCR
- All operations performed under fully traceable Laboratory Information Management System (LIMS)
Production Quality & Proficiency
- Quality assessment, control and characterisation of biospecimens and derivatives (internal and external) applying ISO-17025 accredited methods
- Provision of reference material for biorefinery routine processing
- Proficiency testing programme provider: Assessment of participant compliance with international standards and good practices – FIND OUT MORE.
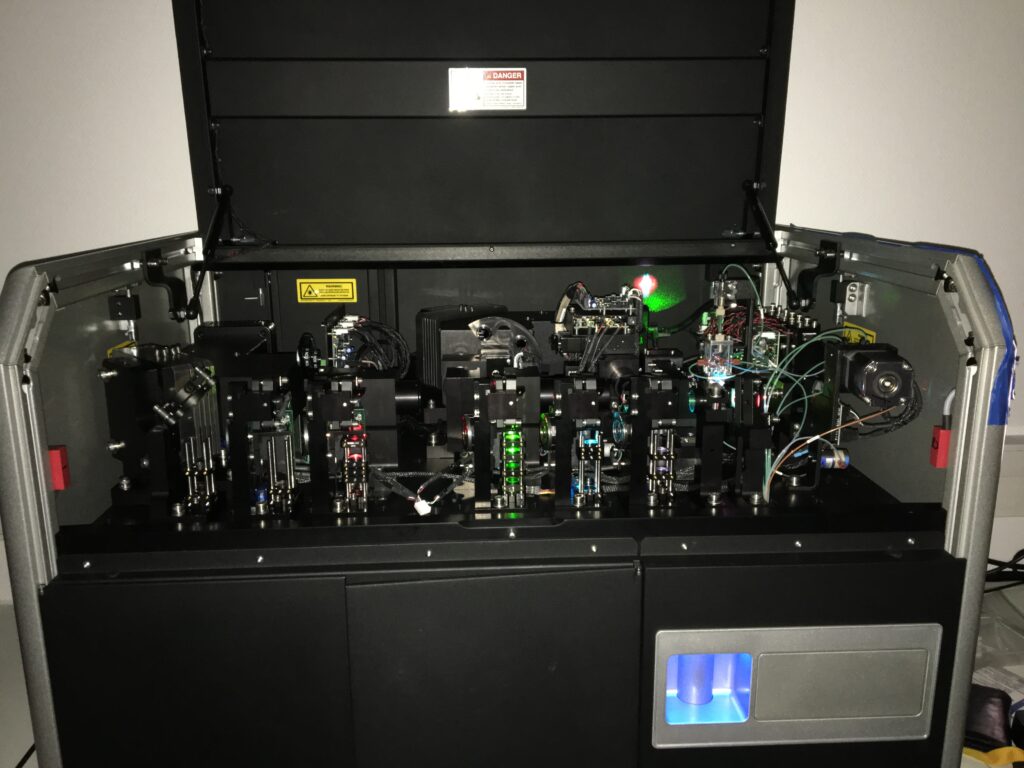
National cytometry platform (iso 9001:2015 certification)
- Technological advice, experimental design and training for flow cytometry, mass cytometry and imaging cytometry and microscopy
- Instrument operations and / or realisation of complete experimental pipelines
- Multidimensional data analysis from sample analysis to data integration and statistics

pathology team
- Histological processing of tissue samples
- Digitalisation of microscopic images of samples
- Pathological characterisation of tissue samples
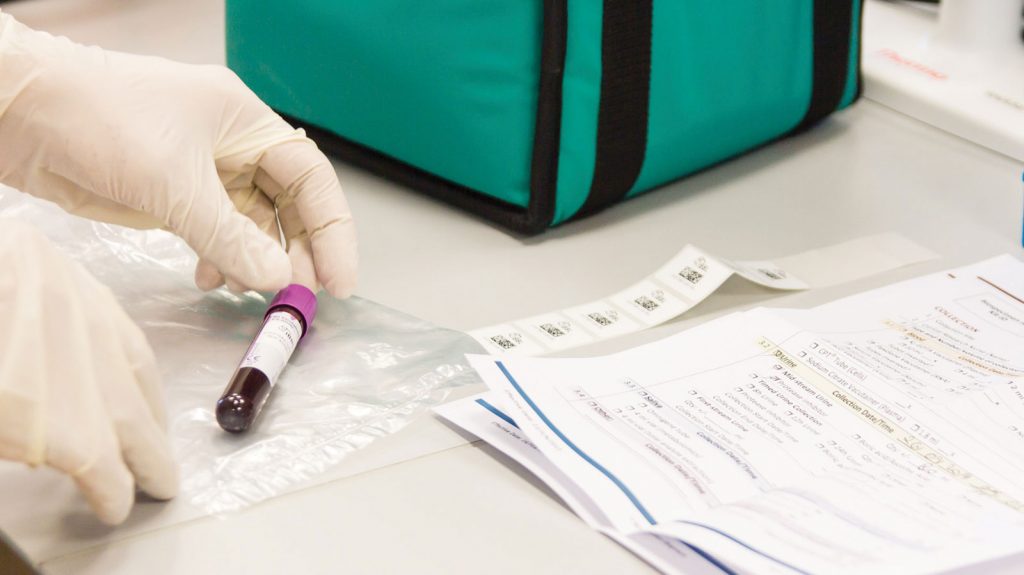
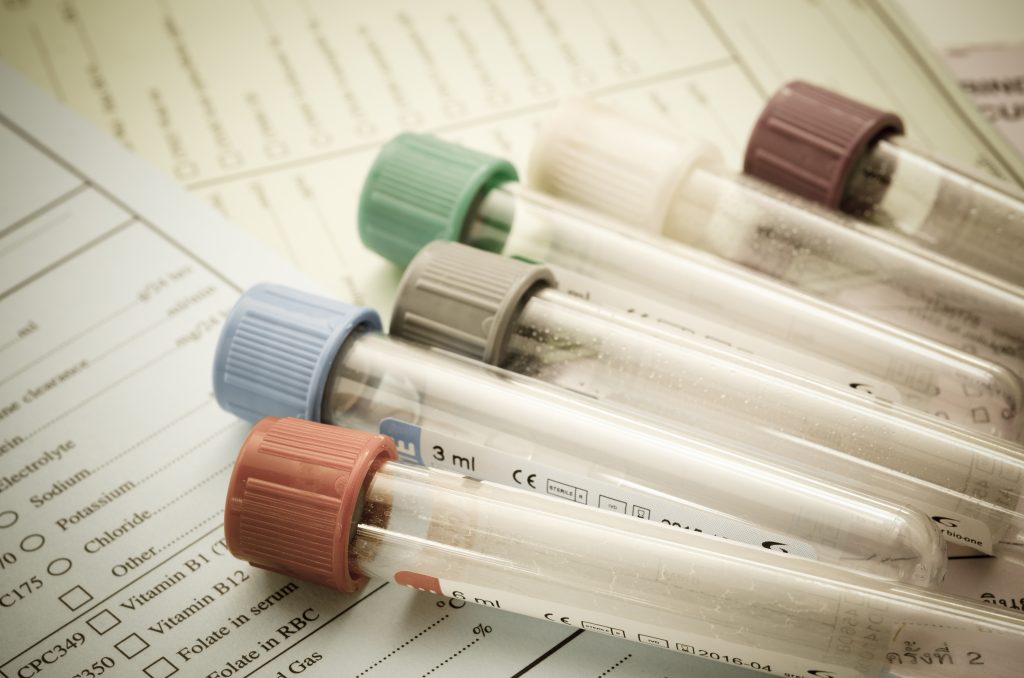
Biorepository (ISO 9001:2015 certification)
As an accredited and certified biobank we operate under a strict quality management system and all our sample logistics and storage activities are carried out by experienced staff following standard operating procedures. We are equipped with 24/7 temperature monitoring and on-call staff, alarm systems, back-up freezers and generators, ensuring the security of your samples.
- Study kit design & provision
- Logistics of biological samples and study related goods in compliance with IATA regulations
- Biological sample receipt, inventory and storage
- Storage equipment and related environmental monitoring management
Translational biomarker group
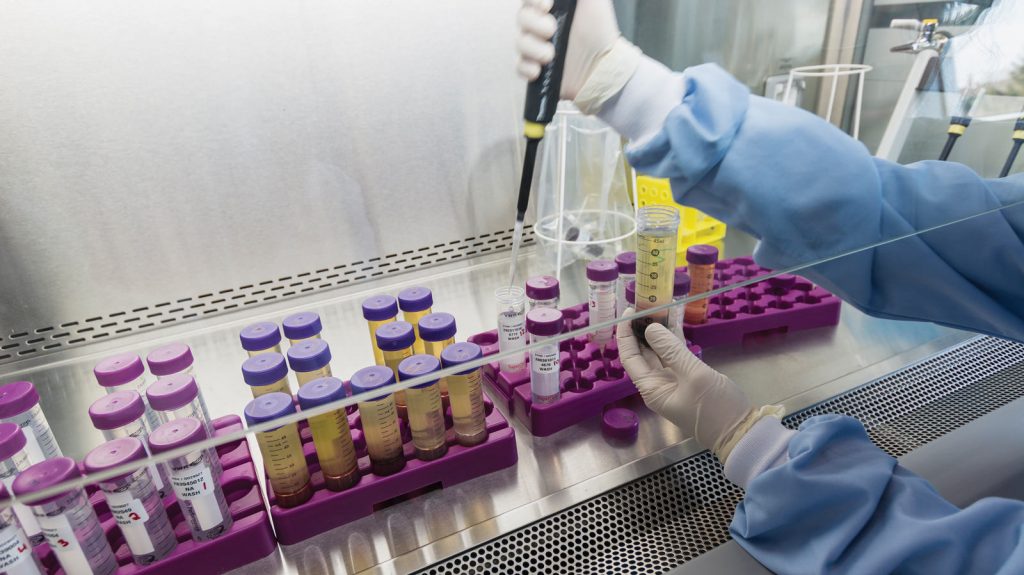
Full-fledged or partial technical biomarker validation, encompassing three modules:
- pre-analytical validation, to assess the most common pre-analytical variables undermining biomarker robustness
- analytical validation, to verify method fitness-for-purpose
- clinical verification, to perform a clinical validation on a smaller scale
Each module can be performed individually or in combination with another module.
Module sequence can be exchanged on a case-by-case BM scenario
Consultancy services for studies revolving around biomarker discovery, also in the context of grant applications Biomarker measurements, including platform bridging (ex.: qPCR to ddrPCR, ELISA to capillary Western Blot, etc.)
Projects & clinical trials
Featured team members
Scientific publications
-
Publisher Correction – 02/09/2022
-
Age at onset as stratifier in idiopathic Parkinson’s disease – effect of ageing and polygenic risk score on clinical phenotypes – 09/08/2022
-
The gut microbial metabolite formate exacerbates colorectal cancer progression – 01/01/2022
-
Results from EDIFICE : A French pilot study on COVID-19 and the gut microbiome in a hospital environment – 08/02/2022
-
Fitness for purpose of stabilized stool samples for bile acid metabolite analyses – 12/04/2021
-
Fibroblast mitochondria in idiopathic Parkinson’s disease display morphological changes and enhanced resistance to depolarization – 01/12/2020
-
Method Validation for Extraction of DNA from Human Stool Samples for Downstream Microbiome Analysis – 01/04/2020
-
Effect of Different Proteinase K Digest Protocols and Deparaffinization Methods on Yield and Integrity of DNA Extracted From Formalin-fixed, Paraffin-embedded Tissue – 01/03/2020
-
Standardization of the preanalytical phase of DNA extraction from fixed tissue for next-generation sequencing analyses – 25/01/2020
-
Blood DNA Yield but Not Integrity or Methylation Is Impacted after Long-Term Storage – 01/01/2016
Related News

Job vacancies
There are no jobs matching this page at the moment. You can view all jobs via the button below.